I cringe when I see research proposals that describe a sophisticated statistical approach, yet do not evaluate this approach in their power/precision/sample size planning. It's often the case that a simplified version of the proposed statistical approach is used instead. Presumably, this is due to the limited availability of power/precision/sample size planning software for sophisticated statistical analyses.
In my own planning, I have defaulted to implementing power/precision analyses with Monte Carlo methods (i.e., simulation). I refer to the approach as "simulated power/precision analysis", but I concede that this may not be the best name. Indeed, there may be a more established name that is unknown to me. This approach initially requires more effort than using one of the many power/precision software packages. However, it's almost always more relevant to the proposed research. With practice, the simulation approach has become second nature, and I use it for complex and simple statistical strategies alike.
Below is an example of the simulation approach to compute the power of a test in a simple crossover design. Whenever a simulated power analysis is implemented, it's necessary to specify (1) how the data will arise, and (2) what statistical procedure will be applied. Note that there is no requirement that the statistical procedure should "match" the data generating mechanism. Rather, it's important that (1) is an accurate reflection of prior belief, and (2) is an accurate representation of the proposed statistical procedure. When (1) and (2) do match, as they do in this example, I am sometimes concerned that the resulting computations are optimistic.
In this example, 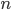 patients will be recruited and given each of two treatments, where the order of treatments is randomized in a block fashion, so that the design is balanced in this regard. We assume that the data arise from a linear mixed effects model, where there is a random intercept for each patient, a treatment effect, an order effect, and a treatment-order interaction effect. The magnitude of each effect is specified, but may be zero. The statistical procedure is to fit a linear mixed effects model, compute a
patients will be recruited and given each of two treatments, where the order of treatments is randomized in a block fashion, so that the design is balanced in this regard. We assume that the data arise from a linear mixed effects model, where there is a random intercept for each patient, a treatment effect, an order effect, and a treatment-order interaction effect. The magnitude of each effect is specified, but may be zero. The statistical procedure is to fit a linear mixed effects model, compute a 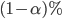 confidence interval for the magnitude of the treatment effect, and finally to make an inference about its significance. We conclude that the treatment effect is (level
confidence interval for the magnitude of the treatment effect, and finally to make an inference about its significance. We conclude that the treatment effect is (level 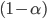 ) significant when the associated confidence interval fails to include the value zero:
) significant when the associated confidence interval fails to include the value zero:
# Simulate a crossover design with the formula:
# Response ~ 1 + Treatment + Order + Treatment:Order + (1 | Patient)
# Fit simulated data with linear mixed effects model. Make
# significance decision about treatment effect on the basis
# of 95% confidence interval (i.e., significant if 95% CI fails
# to include zero).
# n - number of patients in each order group
# sdW - within patient standard deviation
# sdB - between patient standard deviation
# beta - coefficient vector c(Intercept, Treatment, Order, Treatment:Order)
simulate <- function(n, sdW=4, sdB=1, beta=c(8, 4, 0, 0), alpha=0.05) {
require("lme4")
Patient <- as.factor(rep(1:(2*n), rep(2, 2*n)))
Treatment <- c(rep(c("Treatment1", "Treatment2"), n),
rep(c("Treatment2", "Treatment1"), n))
Order <- rep(c("First", "Second"), 2*n)
Data <- data.frame(Patient, Treatment, Order)
FMat <- model.matrix(~ Treatment * Order, data=Data)
RMat <- model.matrix(~ 0 + Patient, data=Data)
Response <- FMat %*% beta + RMat %*% rnorm(2*n, 0, sdB) + rnorm(4*n, 0, sdW)
Data$Response <- Response
Fit <- lmer(Response ~ (1 | Patient) + Treatment * Order, data=Data)
Est <- fixef(Fit)[2]
Ste <- sqrt(vcov(Fit)[2,2])
prod(Est + c(-1,1) * qnorm(1-alpha/2) * Ste) > 0
}
# type I error for n=20 (result: 0.059)
#mean(replicate(1000, simulate(n=20, beta=c(8, 0, 0, 0))))
# type I error for n=50 (result: 0.057)
#mean(replicate(1000, simulate(n=50, beta=c(8, 0, 0, 0))))
# type I error for n=20 and order effect 2 (result: 0.062)
#mean(replicate(1000, simulate(n=20, beta=c(8, 0, 2, 0))))
# type I error for n=50 and order effect 2 (result: 0.05)
#mean(replicate(1000, simulate(n=50, beta=c(8, 0, 2, 0))))
# power for n=20 and treatment effect 4 (result: 0.869)
#mean(replicate(1000, simulate(n=20, beta=c(8, 4, 0, 0))))
# power for n=50 and treatment effect 4 (result: 0.997)
#mean(replicate(1000, simulate(n=50, beta=c(8, 4, 0, 0))))
Several scenarios are considered, including some checks on the type I error associated with the proposed procedure, and its power under three hypothetical data generating mechanisms. ***update 2012/02/23: commenter Paul rightly points out below that 1000 replications is insufficient for the implied precision of three decimal places!*** It's quite late as I'm writing this, and so I will end the discussion here. Indeed, I am trying to shorten my posts in an effort to make them more frequent! Please do comment if I've left out an important detail!
Matt -
You are spot on... when required, power/precision analysis should be completed using simulations approximating the data generation process and the actual analysis. I argue that it should be done even if a "plug-n-play" formula exists in a software package; use the formula to verify the simulation results.
Not that I expect you to program everything for me, but you did ask for comments if details were left out:.
1. Remember to use a seed. Simulations and subsequent results cannot be reproduced without setting a seed.
2. I prefer to simulate through an entire range of plausible effect sizes, saving and plotting both mean and standard error of the simulation at each size.
Thanks Chris. Both of your suggestions are excellent.
Great post! (and not too long, in my book!)
Thanks a ton, I just happened to be looking for something like this. Very helpful.
I hate to point this out, but your replicate size is inadequate for your claimed level of precision. For example three separate runs of your first simulation,
# type I error for n=20 (result: 0.059)
mean(replicate(1000, simulate(n=20, beta=c(8, 0, 0, 0))))
yield results of 0.051, 0.046, 0.06. Clearly, the stated point estimate of 0.059 is a less than accurate representation of the actual precision.
I would suggest that you need to increase the number of replications to 10,000 for two decimal places of accuracy and to 1,000,000 for three decimal places of accuracy, which is what your simulations implicitly claim.
For a paper discussing some simulation size issues see Burton et al, Statist. Med. 2006; 25 : 4279–4292.
Paul, that is an excellent point. Indeed, the items listed in the code block are just for illustration; there is significant Monte Carlo error. Thanks for the citation.